Posted on: 14 March 2022 purlPURL: https://gxy.io/GTN:N00033
We are proud to announce that, as result of the collaboration with the Vertebrate Genomes Project (VGP), a new training describing the VGP assembly pipeline is now available in the Galaxy Training Network. The Vertebrate Genomes Project aims to generate high-quality, near-error-free, gap-free, chromosome-level, haplotype-phased, annotated reference genome assemblies for every vertebrate species.
 Open image in new tab
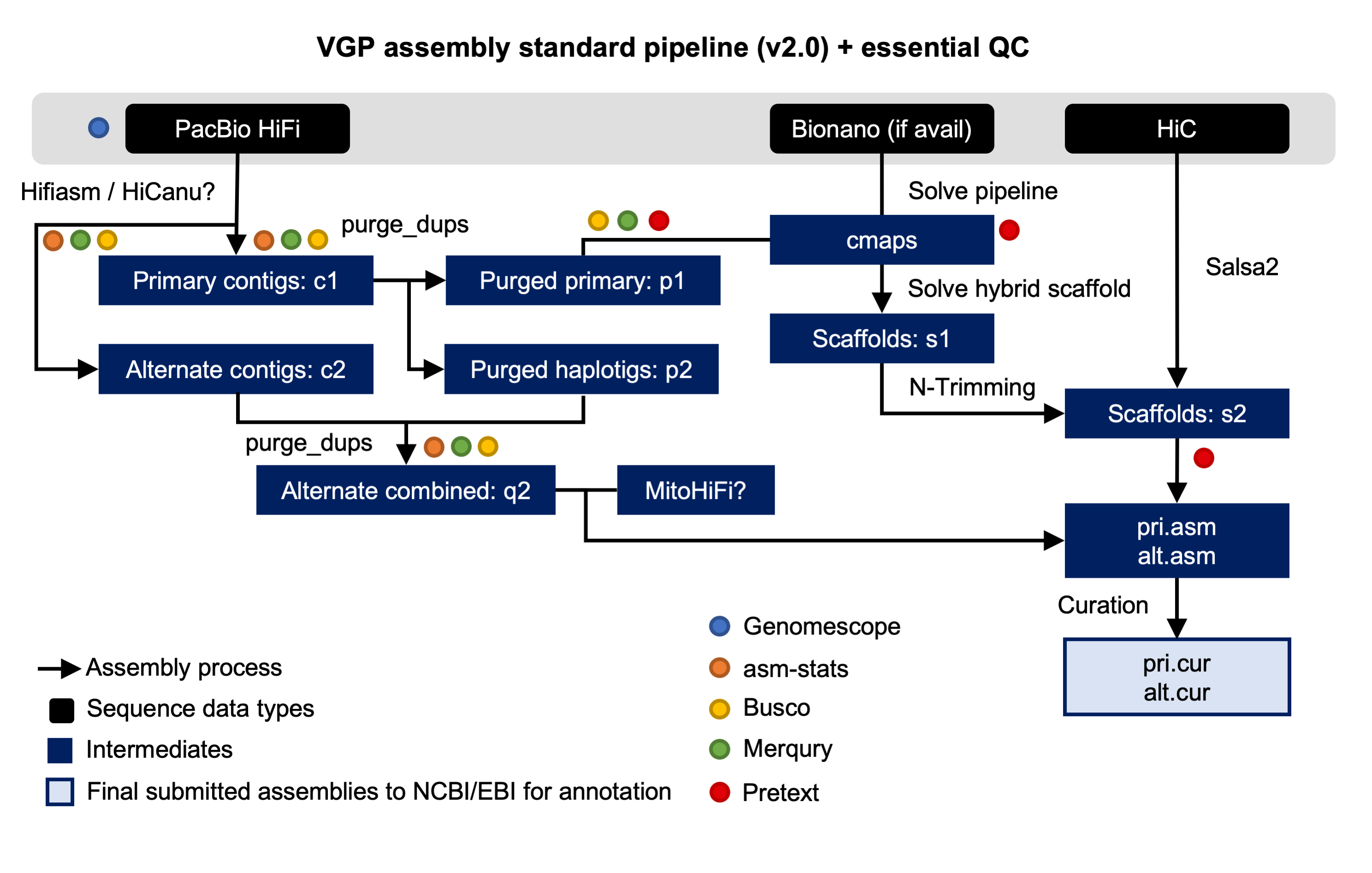
Open image in new tabThe tutorial organized in four sections: genome profile, HiFi phased assembly, post-assembly pocessing and hybrid scaffolding. During the genome profiling stage, diverse tools based on the analsys of k-mer frequencies are used for infering the properties of the genome. After that, a draft assembly is generated by using high accuracy long-read PacBio HiFi reads. In the third stage, the initial assembly is preprocessed for identifying and reassign allelic contigs. Finally, in the last step the assembed contigs are assembled into scaffolds by using two additional technologies: Bionano optical maps and Hi-C data.
View Material