Saskia Hiltemann
Researcher at Erasmus Medical Center
Affiliations
Former Affiliations
Contributions
The following list includes only slides and tutorials where the individual or organisation has been added to the contributor list. This may not include the sum total of their contributions to the training materials (e.g. GTN css or design, tutorial datasets, workflow development, etc.) unless described by a news post.
Editorial Roles
This contributor has taken on additional responsibilities as an editor for the following topics. They are responsible for ensuring that the content is up to date, accurate, and follows GTN best practices.
- Topic: Foundations of Data Science
- Topic: Microbiome
- Topic: Plants
- Topic: Visualisation
- Learning Pathway: Introduction to Galaxy and Sequence analysis
- Learning Pathway: Gallantries Grant - Intellectual Output 1 - Introduction to data analysis and -management, statistics, and coding
- Learning Pathway: Gallantries Grant - Intellectual Output 2 - Large-scale data analysis, and introduction to visualisation and data modelling
- Learning Pathway: Gallantries Grant - Intellectual Output 3 - Data stewardship, federation, standardisation, and collaboration
- Learning Pathway: Creation of a Community CoDex and lab (and optionally of a Special Interest Group (SIG))
Tutorials
- Genome Annotation / Genome Annotation 🧐
- Genome Annotation / Comparative gene analysis in unannotated genomes 🧐
- Genome Annotation / Genome annotation with Braker3 🧐
- Genome Annotation / CRISPR screen analysis 🧐
- Genome Annotation / Secondary metabolite discovery 🧐
- Genome Annotation / Essential genes detection with Transposon insertion sequencing 🧐
- Genome Annotation / Genome annotation with Prokka 🧐
- Genome Annotation / Creating an Official Gene Set 🧐
- Genome Annotation / Masking repeats with RepeatMasker 🧐
- Genome Annotation / Refining Genome Annotations with Apollo (eukaryotes) 🧐
- Genome Annotation / Genome annotation with Helixer 🧐
- Genome Annotation / Genome annotation with Funannotate 🧐
- Genome Annotation / Long non-coding RNAs (lncRNAs) annotation with FEELnc 🧐
- Genome Annotation / Genome annotation with Maker 🧐
- Genome Annotation / From small to large-scale genome comparison 🧐
- Genome Annotation / Refining Genome Annotations with Apollo (prokaryotes) 🧐
- Genome Annotation / Identification of AMR genes in an assembled bacterial genome 📝 🧐
- Genome Annotation / Functional annotation of protein sequences 🧐
- Evolution / Tree thinking for tuberculosis evolution and epidemiology 🧐
- Evolution / Identifying tuberculosis transmission links: from SNPs to transmission clusters 🧐
- Introduction to Galaxy Analyses / Upload data to Galaxy 🧐
- Introduction to Galaxy Analyses / Introduction to Galaxy as an RDM platform ✍️ 🧐
- Introduction to Galaxy Analyses / How to reproduce published Galaxy analyses 🧐
- Introduction to Galaxy Analyses / Best Practices for Citing Galaxy 🧐
- Introduction to Galaxy Analyses / NGS data logistics 🧐
- Introduction to Galaxy Analyses / Very Short Introductions: QC 🧐
- Introduction to Galaxy Analyses / Introduction to Genomics and Galaxy 🧐
- Introduction to Galaxy Analyses / Galaxy Basics for everyone ✍️ 🧐
- Introduction to Galaxy Analyses / Data Manipulation Olympics ✍️ 🧐
- Introduction to Galaxy Analyses / A short introduction to Galaxy 🧐
- Introduction to Galaxy Analyses / IGV Introduction 🧐
- Introduction to Galaxy Analyses / From peaks to genes 🧐
- Introduction to Galaxy Analyses / Galaxy Basics for genomics ✍️ 🧐
- Microbiome / Building an amplicon sequence variant (ASV) table from 16S data using DADA2 🧐
- Microbiome / Calculating α and β diversity from microbiome taxonomic data 🧐
- Microbiome / 16S Microbial Analysis with mothur (short) ✍️ 🧐
- Microbiome / Taxonomic Profiling and Visualization of Metagenomic Data 🧐
- Microbiome / Pathogen detection from (direct Nanopore) sequencing data using Galaxy - Foodborne Edition 🧐
- Microbiome / 16S Microbial analysis with Nanopore data 🧐
- Microbiome / Identification of the micro-organisms in a beer using Nanopore sequencing 🧐
- Microbiome / Antibiotic resistance detection ✍️ 🧐
- Microbiome / MGnify v5.0 Amplicon Pipeline 🧐
- Microbiome / Analyses of metagenomics data - The global picture ✍️ 🧐
- Microbiome / QIIME 2 Moving Pictures 🧐
- Microbiome / Assembly of metagenomic sequencing data 🧐
- Microbiome / QIIME 2 Cancer Microbiome Intervention 🧐
- Microbiome / Metatranscriptomics analysis using microbiome RNA-seq data ✍️ 🧐
- Microbiome / Identifying Mycorrhizal Fungi from ITS2 sequencing using LotuS2 🧐
- Microbiome / 16S Microbial Analysis with mothur (extended) ✍️ 🧐
- Microbiome / Binning of metagenomic sequencing data 🧐
- Microbiome / Metatranscriptomics analysis using microbiome RNA-seq data (short) ✍️ 🧐
- Teaching and Hosting Galaxy training / Teaching experiences 🧐
- Teaching and Hosting Galaxy training / Training techniques to enhance learner participation and engagement 🧐
- Teaching and Hosting Galaxy training / Live Coding is a Skill 🧐
- Teaching and Hosting Galaxy training / Set up a Galaxy for Training ✍️ 🧐
- Teaching and Hosting Galaxy training / Running a workshop as an instructor ✍️ 🧐
- Teaching and Hosting Galaxy training / Course Builder 🧐
- Teaching and Hosting Galaxy training / Motivation and Demotivation 🧐
- Teaching and Hosting Galaxy training / Organizing a workshop 🧐
- Teaching and Hosting Galaxy training / Training Infrastructure as a Service 🧐
- Teaching and Hosting Galaxy training / Train-the-Trainer: putting it all together 🧐
- Teaching and Hosting Galaxy training / Assessment and feedback in training and teachings 🧐
- Teaching and Hosting Galaxy training / Hybrid training 🧐
- Teaching and Hosting Galaxy training / Galaxy Admin Training 🧐
- Teaching and Hosting Galaxy training / Teaching online 🧐
- Teaching and Hosting Galaxy training / Asynchronous training 🧐
- Metabolomics / Mass spectrometry: GC-MS analysis with the metaMS package 🧐
- Metabolomics / Mass spectrometry imaging: Finding differential analytes 🧐
- Metabolomics / Mass spectrometry: LC-MS data processing 🧐
- Metabolomics / Mass spectrometry: GC-MS data processing (with XCMS, RAMClustR, RIAssigner, and matchms) 🧐
- Metabolomics / Mass spectrometry: LC-MS analysis 🧐
- Metabolomics / Mass spectrometry imaging: Examining the spatial distribution of analytes 🧐
- Metabolomics / Predicting EI+ mass spectra with QCxMS 🧐
- Metabolomics / Molecular formula assignment and mass recalibration with MFAssignR package 🧐
- Metabolomics / Mass spectrometry: LC-MS preprocessing with XCMS 🧐
- Single Cell / Generating a single cell matrix using Alevin and combining datasets (bash + R) 🧐
- Single Cell / Understanding Barcodes 🧐
- Single Cell / Importing files from public atlases 🧐
- Single Cell / Downstream Single-cell RNA analysis with RaceID 🧐
- Single Cell / Filter, plot, and explore single cell RNA-seq data with Seurat 🧐
- Single Cell / Removing the effects of the cell cycle 🧐
- Single Cell / Converting between common single cell data formats 🧐
- Single Cell / Inferring single cell trajectories with Scanpy (Python) 🧐
- Single Cell / Bulk RNA Deconvolution with MuSiC 🧐
- Single Cell / Inferring single cell trajectories with Scanpy 🧐
- Single Cell / Converting NCBI Data to the AnnData Format 🧐
- Single Cell / Comparing inferred cell compositions using MuSiC deconvolution 🧐
- Single Cell / Filter, plot and explore single-cell RNA-seq data with Scanpy 🧐
- Single Cell / Inferring single cell trajectories with Monocle3 🧐
- Single Cell / Pseudobulk Analysis with Decoupler and EdgeR 🧐
- Single Cell / Generating a single cell matrix using Alevin 🧐
- Single Cell / Clustering 3K PBMCs with Seurat 📝 🧐
- Single Cell / Single-cell quality control with scater 🧐
- Single Cell / Pre-processing of 10X Single-Cell RNA Datasets 🧐
- Single Cell / GO Enrichment Analysis on Single-Cell RNA-Seq Data 🧐
- Single Cell / Matrix Exchange Format to ESet | Creating a single-cell RNA-seq reference dataset for deconvolution 🧐
- Single Cell / Single-cell ATAC-seq standard processing with SnapATAC2 🧐
- Single Cell / Bulk matrix to ESet | Creating the bulk RNA-seq dataset for deconvolution 🧐
- Single Cell / Clustering 3K PBMCs with Scanpy 🧐
- Single Cell / Inferring single cell trajectories with Monocle3 (R) 🧐
- Single Cell / Evaluating Reference Data for Bulk RNA Deconvolution 🧐
- Single Cell / Filter, plot and explore single-cell RNA-seq data with Scanpy (Python) 🧐
- Single Cell / Scanpy Parameter Iterator 🧐
- Single Cell / Multi-sample batch correction with Harmony and SnapATAC2 🧐
- Single Cell / Combining single cell datasets after pre-processing 🧐
- Single Cell / Pre-processing of Single-Cell RNA Data 🧐
- Single Cell / Filter, plot, and explore single cell RNA-seq data with Seurat (R) 📝 🧐
- Climate / Pangeo ecosystem 101 for everyone - Introduction to Xarray Galaxy Tools 🧐
- Climate / Getting your hands-on earth data 🧐
- Climate / Ocean Data View (ODV) 🧐
- Climate / Analyse Argo data 🧐
- Climate / Collaboration with JupyterGIS 🧐
- Climate / Getting your hands-on climate data 🧐
- Climate / Ocean's variables study 🧐
- Climate / Functionally Assembled Terrestrial Ecosystem Simulator (FATES) with Galaxy Climate JupyterLab 🧐
- Climate / Functionally Assembled Terrestrial Ecosystem Simulator (FATES) 🧐
- Climate / Nitrate DMQC for autonomous platforms such as Argo floats 🧐
- Climate / Visualize Climate data with Panoply netCDF viewer 🧐
- Climate / Pangeo Notebook in Galaxy - Introduction to Xarray 🧐
- Climate / Sentinel 5P data visualisation 🧐
- Synthetic Biology / Designing plasmids encoding predicted pathways by using the BASIC assembly method 🧐
- Synthetic Biology / Evaluating and ranking a set of pathways based on multiple metrics 🧐
- Synthetic Biology / Generating theoretical possible pathways for the production of Lycopene in E.Coli using Retrosynthesis tools 🧐
- Variant Analysis / Trio Analysis using Synthetic Datasets from RD-Connect GPAP ✍️ 🧐
- Variant Analysis / Exome sequencing data analysis for diagnosing a genetic disease 🧐
- Variant Analysis / Calling variants in diploid systems 🧐
- Variant Analysis / Identification of somatic and germline variants from tumor and normal sample pairs 🧐
- Variant Analysis / Working with Beacon V2: A Comprehensive Guide to Creating, Uploading, and Searching for Variants with Beacons 🧐
- Variant Analysis / Somatic Variant Discovery from WES Data Using Control-FREEC 🧐
- Variant Analysis / Calling variants in non-diploid systems 🧐
- Variant Analysis / Avian influenza viral strain analysis from gene segment sequencing data 🧐
- Variant Analysis / Microbial Variant Calling 🧐
- Variant Analysis / Pox virus genome analysis from tiled-amplicon sequencing data 🧐
- Variant Analysis / M. tuberculosis Variant Analysis 🧐
- Variant Analysis / From NCBI's Sequence Read Archive (SRA) to Galaxy: SARS-CoV-2 variant analysis 🧐
- Variant Analysis / Calling very rare variants 🧐
- Variant Analysis / Mutation calling, viral genome reconstruction and lineage/clade assignment from SARS-CoV-2 sequencing data 🧐
- Variant Analysis / Querying the University of Bradford GDC Beacon Database for Copy Number Variants (CNVs) 🧐
- Variant Analysis / Mapping and molecular identification of phenotype-causing mutations 🧐
- Sequence analysis / Removal of human reads from SARS-CoV-2 sequencing data 🧐
- Sequence analysis / Quality and contamination control in bacterial isolate using Illumina MiSeq Data 🧐
- Sequence analysis / NCBI BLAST+ against the MAdLand 🧐
- Sequence analysis / Screening assembled genomes for contamination using NCBI FCS 🧐
- Sequence analysis / Quality Control 🧐
- Sequence analysis / Mapping 🧐
- Sequence analysis / Clean and manage Sanger sequences from raw files to aligned consensus 🧐
- Sequence analysis / Identification and Evolutionary Analysis of Transcription-Associated Proteins in Streptophyte algae and Land plants ✍️ 🧐
- Visualisation / Genomic Data Visualisation with JBrowse ✍️ 🧐
- Visualisation / Visualisation with Circos ✍️ 🧐
- Visualisation / Ploting a Microbial Genome with Circos 🧐
- Galaxy Server administration / Ansible ✍️ 🧐
- Galaxy Server administration / Deploying Wireguard for private mesh networking 🧐
- Galaxy Server administration / Galaxy usage on SURF Research Cloud 🧐
- Galaxy Server administration / Galaxy Interactive Tools 🧐
- Galaxy Server administration / Reference Data with CVMFS 🧐
- Galaxy Server administration / Performant Uploads with TUS 🧐
- Galaxy Server administration / Create a subdomain for your community on UseGalaxy.eu 🧐
- Galaxy Server administration / Galaxy Monitoring with Reports 🧐
- Galaxy Server administration / Upgrading Galaxy 🧐
- Galaxy Server administration / Deploying Tailscale/Headscale for private mesh networking 🧐
- Galaxy Server administration / Setting up Celery Workers for Galaxy 🧐
- Galaxy Server administration / External Authentication 🧐
- Galaxy Server administration / Mapping Jobs to Destinations using TPV 🧐
- Galaxy Server administration / How I learned to stop worrying and love the systemd 🧐
- Galaxy Server administration / Deploying a compute cluster in OpenStack via Terraform 🧐
- Galaxy Server administration / Automation with Jenkins 🧐
- Galaxy Server administration / Managing Galaxy on Kubernetes 🧐
- Galaxy Server administration / Galaxy Tool Management with Ephemeris 🧐
- Galaxy Server administration / Reference Data with CVMFS without Ansible 🧐
- Galaxy Server administration / Server Maintenance: Cleanup, Backup, and Restoration 🧐
- Galaxy Server administration / Deploying a Beacon v1 in Galaxy 📝 🧐
- Galaxy Server administration / Galaxy Installation with Ansible 📝 🧐
- Galaxy Server administration / Monitoring Galaxy and Pulsar with Sentry 🧐
- Galaxy Server administration / Reference Data with Data Managers 🧐
- Galaxy Server administration / Enable upload via FTP 🧐
- Galaxy Server administration / Use Apptainer containers for running Galaxy jobs 🧐
- Galaxy Server administration / Galaxy Monitoring with Telegraf and Grafana 🧐
- Galaxy Server administration / Customizing the look of Galaxy 🧐
- Galaxy Server administration / Pulsar usage on SURF Research Cloud 🧐
- Galaxy Server administration / Galaxy Installation on Kubernetes 🧐
- Galaxy Server administration / Data Libraries ✍️ 🧐
- Galaxy Server administration / Running Jobs on Remote Resources with Pulsar 🧐
- Galaxy Server administration / Galaxy Database schema 🧐
- Galaxy Server administration / Connecting Galaxy to a compute cluster 🧐
- Galaxy Server administration / Galaxy Monitoring with gxadmin 🧐
- Galaxy Server administration / Distributed Object Storage 🧐
- Galaxy Server administration / Training Infrastructure as a Service (TIaaS) ✍️ 🧐
- Proteomics / Machine Learning Modeling of Anticancer Peptides 🧐
- Proteomics / Neoantigen 3: PepQuery2 Verification 🧐
- Proteomics / Clinical Metaproteomics 1: Database-Generation 🧐
- Proteomics / Label-free versus Labelled - How to Choose Your Quantitation Method 🧐
- Proteomics / Clinical Metaproteomics 3: Verification 🧐
- Proteomics / Proteogenomics 3: Novel peptide analysis 🧐
- Proteomics / Mass spectrometry imaging: Loading and exploring MSI data 🧐
- Proteomics / Neoantigen 5a: Predicting HLA Binding 🧐
- Proteomics / metaQuantome 1: Data creation 🧐
- Proteomics / MaxQuant and MSstats for the analysis of label-free data 🧐
- Proteomics / Secretome Prediction 🧐
- Proteomics / metaQuantome 3: Taxonomy 🧐
- Proteomics / Clinical Metaproteomics 4: Quantitation 🧐
- Proteomics / Peptide and Protein ID using SearchGUI and PeptideShaker 🧐
- Proteomics / EncyclopeDIA 🧐
- Proteomics / Metaproteomics tutorial 🧐
- Proteomics / Peptide and Protein Quantification via Stable Isotope Labelling (SIL) 🧐
- Proteomics / Proteogenomics 2: Database Search 🧐
- Proteomics / Detection and quantitation of N-termini (degradomics) via N-TAILS 🧐
- Proteomics / Library Generation for DIA Analysis 🧐
- Proteomics / MaxQuant and MSstats for the analysis of TMT data 🧐
- Proteomics / DIA Analysis using OpenSwathWorkflow 🧐
- Proteomics / Neoantigen 1a: Fusion-Database-Generation 🧐
- Proteomics / Neoantigen 2: Database merge and FragPipe discovery 🧐
- Proteomics / Peptide and Protein ID using OpenMS tools 🧐
- Proteomics / Annotating a protein list identified by LC-MS/MS experiments 🧐
- Proteomics / metaQuantome 2: Function 🧐
- Proteomics / Neoantigen 4: Variant Annotation 🧐
- Proteomics / Protein FASTA Database Handling 🧐
- Proteomics / Peptide Library Data Analysis 🧐
- Proteomics / Clinical Metaproteomics 5: Data Interpretation 🧐
- Proteomics / Biomarker candidate identification 🧐
- Proteomics / Label-free data analysis using MaxQuant 🧐
- Proteomics / Proteogenomics 1: Database Creation 🧐
- Proteomics / Neoantigen 1b: Non-Reference-Database-Generation 🧐
- Proteomics / Neoantigen 5b: IEDB binding Validated Neopeptides 🧐
- Proteomics / Multiomics data analysis using MultiGSEA 🧐
- Proteomics / Clinical Metaproteomics 2: Discovery 🧐
- Digital Humanities / Introduction to Digital Humanities in Galaxy 🧐
- Digital Humanities / OpenRefine Tutorial for researching cultural data 🧐
- Digital Humanities / Text-Mining Differences in Chinese Newspaper Articles 🧐
- Digital Humanities / Transcribing Audio and Video files with Automated Speech Recognition 🧐
- Using Galaxy and Managing your Data / JupyterLab in Galaxy 🧐
- Using Galaxy and Managing your Data / Submitting sequence data to ENA 🧐
- Using Galaxy and Managing your Data / Extracting Workflows from Histories 🧐
- Using Galaxy and Managing your Data / Creating, Editing and Importing Galaxy Workflows 🧐
- Using Galaxy and Managing your Data / Workflow Reports ✍️ 🧐
- Using Galaxy and Managing your Data / InterMine integration with Galaxy 🧐
- Using Galaxy and Managing your Data / Understanding Galaxy history system 📝 🧐
- Using Galaxy and Managing your Data / Use Jupyter notebooks in Galaxy 🧐
- Using Galaxy and Managing your Data / Group tags for complex experimental designs 🧐
- Using Galaxy and Managing your Data / Using Workflow Parameters 🧐
- Using Galaxy and Managing your Data / Rule Based Uploader 🧐
- Using Galaxy and Managing your Data / Automating Galaxy workflows using the command line 🧐
- Using Galaxy and Managing your Data / Using dataset collections 🧐
- Using Galaxy and Managing your Data / Rule Based Uploader: Advanced 🧐
- Using Galaxy and Managing your Data / Name tags for following complex histories 🧐
- Using Galaxy and Managing your Data / Downloading and Deleting Data in Galaxy 🧐
- Using Galaxy and Managing your Data / SRA Aligned Read Format to Speed Up SARS-CoV-2 data Analysis 🧐
- Using Galaxy and Managing your Data / Searching Your History 🧐
- Using Galaxy and Managing your Data / RStudio in Galaxy 🧐
- Computational chemistry / Setting up molecular systems 🧐
- Computational chemistry / Virtual screening of the SARS-CoV-2 main protease with rxDock and pose scoring 🧐
- Computational chemistry / Protein-ligand docking 🧐
- Computational chemistry / High Throughput Molecular Dynamics and Analysis 🧐
- Computational chemistry / Analysis of molecular dynamics simulations 🧐
- Computational chemistry / Protein target prediction of a bioactive ligand with Align-it and ePharmaLib 🧐
- Computational chemistry / Running molecular dynamics simulations using GROMACS 🧐
- Computational chemistry / Running molecular dynamics simulations using NAMD 🧐
- Ecology / Checking expected species and contamination in bacterial isolate 🧐
- Ecology / Data submission using ENA upload Tool 🧐
- Ecology / Marine Omics identifying biosynthetic gene clusters 🧐
- Ecology / Obis marine indicators 🧐
- Ecology / Biodiversity data exploration 🧐
- Ecology / Cleaning GBIF data using OpenRefine 🧐
- Ecology / Taxonomic Analysis of eDNA 🧐
- Ecology / Ecoregionalization workflow tutorial 🧐
- Ecology / Sentinel 2 biodiversity 🧐
- Ecology / Phylodiversity analysis quick tutorial 🧐
- Ecology / RAD-Seq to construct genetic maps 🧐
- Ecology / Species distribution modeling 🧐
- Ecology / Audio data annotation with NEAL (Nature + Energy Audio labeler) 🧐
- Ecology / Visualization of Climate Data using NetCDF xarray Map Plotting 🧐
- Ecology / Compute and analyze biodiversity metrics with PAMPA toolsuite 🧐
- Ecology / Creating metadata using Ecological Metadata Language (EML) standard with EML Assembly Line functionalities 🧐
- Ecology / Regional GAM 🧐
- Ecology / Life Traits Ecoregionalization workflow 🧐
- Ecology / RAD-Seq de-novo data analysis 🧐
- Ecology / Metabarcoding/eDNA through Obitools 🧐
- Ecology / QGIS Web Feature Services 🧐
- Ecology / Visualize EBV cube data with Panoply netCDF viewer 🧐
- Ecology / Creating FAIR Quality assessment reports and draft of Data Papers from EML metadata with MetaShRIMPS 🧐
- Ecology / RAD-Seq Reference-based data analysis 🧐
- Ecology / Preparing genomic data for phylogeny reconstruction 🧐
- Ecology / Champs blocs indicators 🧐
- Ecology / Cleaning GBIF data for the use in Ecology 🧐
- Galaxy Community Building / Creating a Special Interest Group 📝 🧐
- Galaxy Community Building / Make your tools available on your subdomain 🧐
- Galaxy Community Building / Creation of resources listing all the tools and their metadata relevant to your community 🧐
- Galaxy Community Building / What's a Special Interest Group? 📝 🧐
- Galaxy Community Building / Creation of resources listing Galaxy workflow for your community 🧐
- Galaxy Community Building / Creation of a Galaxy tutorial table for your community 🧐
- Galaxy Community Building / Creation of the labs in the different Galaxy instances for your community 🧐
- Galaxy Community Building / Creating community content ✍️ 🧐
- Foundations of Data Science / Python - Coding Style 🧐
- Foundations of Data Science / R basics in Galaxy 🧐
- Foundations of Data Science / Make & Snakemake 🧐
- Foundations of Data Science / Python - Warm-up for statistics and machine learning 🧐
- Foundations of Data Science / Python - Functions 🧐
- Foundations of Data Science / Introduction to sequencing with Python (part two) 🧐
- Foundations of Data Science / Python - Testing 🧐
- Foundations of Data Science / SQL Educational Game - Murder Mystery 🧐
- Foundations of Data Science / Python - Argparse 🧐
- Foundations of Data Science / Python - Files & CSV 🧐
- Foundations of Data Science / Conda Environments For Software Development 🧐
- Foundations of Data Science / Data visualisation Olympics - Visualization in R ⚙️ 🧐
- Foundations of Data Science / Python - Type annotations 🧐
- Foundations of Data Science / Data Manipulation Olympics - SQL ✍️ 🧐
- Foundations of Data Science / Virtual Environments For Software Development 🧐
- Foundations of Data Science / Version Control with Git 📝 🧐
- Foundations of Data Science / Introduction to sequencing with Python (part four) 🧐
- Foundations of Data Science / Python - Introductory Graduation 🧐
- Foundations of Data Science / Variant Calling Workflow ✍️ 🧐
- Foundations of Data Science / Advanced CLI in Galaxy 🧐
- Foundations of Data Science / Introduction to Python 🧐
- Foundations of Data Science / Introduction to SQL 🧐
- Foundations of Data Science / Python - Math 🧐
- Foundations of Data Science / dplyr & tidyverse for data processing 🧐
- Foundations of Data Science / Advanced SQL 🧐
- Foundations of Data Science / One protein along the UniProt page 🧐
- Foundations of Data Science / CLI Educational Game - Bashcrawl 🧐
- Foundations of Data Science / SQL with Python 🧐
- Foundations of Data Science / Data Manipulation Olympics - JQ ✍️ 🧐
- Foundations of Data Science / Python - Multiprocessing 🧐
- Foundations of Data Science / Python - Lists & Strings & Dictionaries 🧐
- Foundations of Data Science / Advanced Python 🧐
- Foundations of Data Science / Learning about one gene across biological resources and formats 🧐
- Foundations of Data Science / Data manipulation with Pandas 🧐
- Foundations of Data Science / A (very) brief history of genomics 🧐
- Foundations of Data Science / Plotting in Python 🧐
- Foundations of Data Science / CLI basics 🧐
- Foundations of Data Science / Basics of using Git from the Command Line 🧐
- Foundations of Data Science / SQL with R 🧐
- Foundations of Data Science / Advanced R in Galaxy 🧐
- Imaging / Quantification of electrophoresis gel bands using QuPath and Galaxy imaging tools 🧐
- Imaging / End-to-End Tissue Microarray Image Analysis with Galaxy-ME 🧐
- Imaging / Introduction to Image Analysis using Galaxy ✍️ 🧐
- Imaging / Tracking of mitochondria and capturing mitoflashes 🧐
- Imaging / Overview of the Galaxy OMERO-suite - Upload images and metadata in OMERO using Galaxy 🧐
- Imaging / Object tracking using CellProfiler 🧐
- Imaging / Segmentation of Anatomical Structures in Medical 3-D Images 🧐
- Imaging / Voronoi segmentation 🧐
- Imaging / Quantification of single-molecule RNA fluorescence in situ hybridization (smFISH) in yeast cell lines 🧐
- Imaging / Execute a BiaPy workflow in Galaxy 🧐
- Imaging / Training Custom YOLO Models for Segmentation of Bioimages 🧐
- Imaging / Using BioImage.IO models for image analysis in Galaxy 🧐
- Imaging / Analyse HeLa fluorescence siRNA screen 🧐
- Imaging / Nucleoli segmentation and feature extraction using CellProfiler 🧐
- Imaging / Parameter tuning and optimization - Evaluating nuclei segmentation with Galaxy 🧐
- Transcriptomics / CLIP-Seq data analysis from pre-processing to motif detection 🧐
- Transcriptomics / Whole transcriptome analysis of Arabidopsis thaliana 🧐
- Transcriptomics / Reference-based RNAseq data analysis (long) 🧐
- Transcriptomics / RNA-Seq data analysis, clustering and visualisation tutorial 📝 🧐
- Transcriptomics / De novo transcriptome reconstruction with RNA-Seq 🧐
- Transcriptomics / Differential abundance testing of small RNAs 🧐
- Transcriptomics / RNA-RNA interactome data analysis 🧐
- Transcriptomics / Reference-based RNA-Seq data analysis 🧐
- Transcriptomics / 1: RNA-Seq reads to counts 🧐
- Transcriptomics / Visualization of RNA-Seq results with heatmap2 🧐
- Transcriptomics / RNA-Seq analysis with AskOmics Interactive Tool 🧐
- Transcriptomics / Visualization of RNA-Seq results with Volcano Plot in R 🧐
- Transcriptomics / Pathway analysis with the MINERVA Platform ✍️ 🧐
- Transcriptomics / Network analysis with Heinz 🧐
- Transcriptomics / Genome-wide alternative splicing analysis 🧐
- Transcriptomics / 2: RNA-seq counts to genes 🧐
- Transcriptomics / Visualization of RNA-Seq results with Volcano Plot 🧐
- Transcriptomics / GO Enrichment Analysis 🧐
- Transcriptomics / RNA Seq Counts to Viz in R 🧐
- Transcriptomics / RNA-seq Alignment with STAR 📝 🧐
- Transcriptomics / Small Non-coding RNA Clustering using BlockClust 🧐
- Transcriptomics / Visualization of RNA-Seq results with CummeRbund 🧐
- Transcriptomics / De novo transcriptome assembly, annotation, and differential expression analysis 🧐
- Epigenetics / CUT&RUN data analysis 🧐
- Epigenetics / Infinium Human Methylation BeadChip 🧐
- Epigenetics / Hi-C analysis of Drosophila melanogaster cells using HiCExplorer 🧐
- Epigenetics / Identification of the binding sites of the T-cell acute lymphocytic leukemia protein 1 (TAL1) 🧐
- Epigenetics / DNA Methylation data analysis 🧐
- Epigenetics / Formation of the Super-Structures on the Inactive X 🧐
- Epigenetics / Identification of the binding sites of the Estrogen receptor 🧐
- Epigenetics / ATAC-Seq data analysis 🧐
- Contributing to the Galaxy Training Material / Creating Interactive Galaxy Tours ✍️ 🧐
- Contributing to the Galaxy Training Material / Teaching Python 🧐
- Contributing to the Galaxy Training Material / FAIR-by-Design methodology 🧐
- Contributing to the Galaxy Training Material / Including a new topic 🧐
- Contributing to the Galaxy Training Material / Contributing to the Galaxy Training Network with GitHub 🧐
- Contributing to the Galaxy Training Material / Creating a new tutorial ✍️ 🧐
- Contributing to the Galaxy Training Material / Design and plan session, course, materials 🧐
- Contributing to the Galaxy Training Material / Contributing with GitHub via its interface 🧐
- Contributing to the Galaxy Training Material / Tools, Data, and Workflows for tutorials ✍️ 🧐
- Contributing to the Galaxy Training Material / Preview the GTN website as you edit your training material ✍️ 🧐
- Contributing to the Galaxy Training Material / Principles of learning and how they apply to training and teaching 🧐
- Contributing to the Galaxy Training Material / Generating PDF artefacts of the website 🧐
- Contributing to the Galaxy Training Material / Updating diffs in admin training 🧐
- Contributing to the Galaxy Training Material / Adding auto-generated video to your slides 🧐
- Contributing to the Galaxy Training Material / GTN Metadata 🧐
- Contributing to the Galaxy Training Material / Creating content in Markdown 📝 🧐
- Contributing to the Galaxy Training Material / Adding Quizzes to your Tutorial 🧐
- FAIR Data, Workflows, and Research / Submitting workflows to LifeMonitor 🧐
- FAIR Data, Workflows, and Research / Uploading Data to Zenodo from Galaxy 🧐
- FAIR Data, Workflows, and Research / DataPLANT ARCs ✍️ 🧐
- FAIR Data, Workflows, and Research / Best practices for workflows in GitHub repositories 🧐
- FAIR Data, Workflows, and Research / FAIR Bioimage Metadata 🧐
- FAIR Data, Workflows, and Research / FAIR data management solutions 🧐
- FAIR Data, Workflows, and Research / Integrating InvenioRDM-compatible Repositories with Galaxy 🧐
- FAIR Data, Workflows, and Research / Data Registration 🧐
- FAIR Data, Workflows, and Research / Workflow Run RO-Crate Introduction 🧐
- FAIR Data, Workflows, and Research / FAIR Galaxy Training Material 🧐
- FAIR Data, Workflows, and Research / Sequence data submission to ENA 🧐
- FAIR Data, Workflows, and Research / REMBI - Recommended Metadata for Biological Images – metadata guidelines for bioimaging data 🧐
- FAIR Data, Workflows, and Research / RO-Crate - Introduction 🧐
- FAIR Data, Workflows, and Research / FAIR in a nutshell 🧐
- FAIR Data, Workflows, and Research / Exporting Workflow Run RO-Crates from Galaxy 🧐
- FAIR Data, Workflows, and Research / RO-Crate in Python 🧐
- FAIR Data, Workflows, and Research / Introduction to Data Management Plan (DMP) for Peatland Research and PeatDataHub 🧐
- FAIR Data, Workflows, and Research / Making clinical datasets FAIR 🧐
- Statistics and machine learning / GLEAM Image Learner - Validating Skin Lesion Classification on HAM10000 🧐
- Statistics and machine learning / Text-mining with the SimText toolset 🧐
- Statistics and machine learning / Foundational Aspects of Machine Learning using Python 🧐
- Statistics and machine learning / Deep Learning (Part 3) - Convolutional neural networks (CNN) 🧐
- Statistics and machine learning / Interval-Wise Testing for omics data 🧐
- Statistics and machine learning / Basics of machine learning 🧐
- Statistics and machine learning / Classification in Machine Learning 🧐
- Statistics and machine learning / Fine tune large protein model (ProtTrans) using HuggingFace 🧐
- Statistics and machine learning / Gleam Multimodal Learner - Head and Neck cancer Recurrence Prediction with HANCOCK 🧐
- Statistics and machine learning / Train and Test a Deep learning image classifier with Galaxy-Ludwig 🧐
- Statistics and machine learning / PAPAA PI3K_OG: PanCancer Aberrant Pathway Activity Analysis 🧐
- Statistics and machine learning / Age prediction using machine learning 🧐
- Statistics and machine learning / Introduction to Machine Learning using R 🧐
- Statistics and machine learning / Clustering in Machine Learning 🧐
- Statistics and machine learning / Deep Learning (Part 2) - Recurrent neural networks (RNN) 🧐
- Statistics and machine learning / Image classification in Galaxy with fruit 360 dataset 🧐
- Statistics and machine learning / Neural networks using Python 🧐
- Statistics and machine learning / Introduction to deep learning 🧐
- Statistics and machine learning / Deep Learning (Part 1) - Feedforward neural networks (FNN) 🧐
- Statistics and machine learning / Deep Learning (without Generative Artificial Intelligence) using Python 🧐
- Statistics and machine learning / Regulations/standards for AI using DOME 🧐
- Statistics and machine learning / A Docker-based interactive Jupyterlab powered by GPU for artificial intelligence in Galaxy 🧐
- Statistics and machine learning / Machine learning: classification and regression 🧐
- Statistics and machine learning / Regression in Machine Learning 🧐
- Statistics and machine learning / Galaxy Tabular Learner - Building a Model using Chowell clinical data 🧐
- Assembly / Genome assembly using PacBio data 🧐
- Assembly / Using the VGP workflows to assemble a vertebrate genome with HiFi and Hi-C data 🧐
- Assembly / Unicycler assembly of SARS-CoV-2 genome with preprocessing to remove human genome reads 🧐
- Assembly / An Introduction to Genome Assembly 🧐
- Assembly / Vertebrate genome assembly using HiFi, Bionano and Hi-C data - Step by Step 🧐
- Assembly / Decontamination of a genome assembly 🧐
- Assembly / Genome Assembly Quality Control 🧐
- Assembly / Unicycler Assembly 🧐
- Assembly / Making sense of a newly assembled genome 🧐
- Assembly / Large genome assembly and polishing 🧐
- Assembly / De Bruijn Graph Assembly ✍️ 🧐
- Assembly / Chloroplast genome assembly 🧐
- Assembly / ERGA post-assembly QC 🧐
- Assembly / Genome Assembly of MRSA from Oxford Nanopore MinION data (and optionally Illumina data) 📝 🧐
- Assembly / Genome Assembly of a bacterial genome (MRSA) sequenced using Illumina MiSeq Data 📝 🧐
- Development in Galaxy / Galaxy Interactive Tools 🧐
- Development in Galaxy / Creating Galaxy tools from Conda Through Deployment 🧐
- Development in Galaxy / ToolFactory: Generating Tools From Simple Scripts 🧐
- Development in Galaxy / Galaxy Webhooks 🧐
- Development in Galaxy / Contributing to BioBlend as a developer 🧐
- Development in Galaxy / nb2workflow: Generating Galaxy Tools From Jupyter Notebooks 🧐
- Development in Galaxy / Contributing a New Feature to Galaxy Core 🧐
- Development in Galaxy / Debugging Galaxy 🧐
- Development in Galaxy / Writing Automated Tests for Galaxy 🧐
- Development in Galaxy / Scripting Galaxy using the API and BioBlend 🧐
- Development in Galaxy / Adding and updating best practice metadata for Galaxy tools using the bio.tools registry 🧐
- Development in Galaxy / Generic plugins ✍️ 🧐
- Development in Galaxy / ToolFactory: Generating Tools From More Complex Scripts 🧐
- Development in Galaxy / JavaScript plugins ✍️ 🧐
- Development in Galaxy / Data source integration ✍️ 🧐
- Introduction to Galaxy Analyses / Von Peaks zu Genen 🧐
- Sequence analysis / Qualitätskontrolle 🧐
- Sequence analysis / Mapping 🧐
- Foundations of Data Science / Ein Protein entlang der UniProt-Seite 🧐
- Foundations of Data Science / Lernen über ein Gen über biologische Ressourcen und Formate hinweg 🧐
- Transcriptomics / Referenzbasierte RNA-Seq-Datenanalyse 🧐
- Introduction to Galaxy Analyses / Breve introducción a Galaxy - en español 🧐
- Introduction to Galaxy Analyses / De picos a genes 🧐
- Sequence analysis / Control de calidad 🧐
- Sequence analysis / Mapeo 🧐
- Foundations of Data Science / Una proteína a lo largo de la página UniProt 🧐
- Foundations of Data Science / Aprendizaje sobre un gen a través de recursos y formatos biológicos 🧐
- Transcriptomics / Análisis de datos RNA-Seq basados en referencias 🧐
- Ecology / Production d'indicateurs champs de bloc 🧐
- Introduction to Galaxy Analyses / Dai picchi ai geni 🧐
- Sequence analysis / Controllo qualità 🧐
- Sequence analysis / Mappatura 🧐
- Foundations of Data Science / Una proteina lungo la pagina UniProt 🧐
- Foundations of Data Science / Imparare a conoscere un gene attraverso risorse e formati di dato biologici 🧐
- Transcriptomics / Analisi dei dati RNA-Seq basata su riferimenti 🧐
Slides
- Synthetic Biology / Introduction to Synthetic Biology 🧐
- Development in Galaxy / Galaxy from a developer point of view 🧐
- Genome Annotation / Introduction to Genome Annotation 🧐
- Genome Annotation / Genome annotation with Prokka 🧐
- Genome Annotation / High Performance Computing for Pairwise Genome Comparison 🧐
- Genome Annotation / Refining Genome Annotations with Apollo 🧐
- Evolution / Phylogenetics - Back to Basics - Introduction 🧐
- Evolution / Phylogenetics - Back to Basics - Multiple Sequence Alignment 🧐
- Evolution / Phylogenetics - Back to Basics - Estimating trees from alignments 🧐
- Evolution / Phylogenetics - Back to Basics - Phylogenetic Networks 🧐
- Evolution / Phylogenetics - Back to Basics - Terminology 🧐
- Evolution / Phylogenetics - Back to Basics - Building Trees 🧐
- Introduction to Galaxy Analyses / Introduction to Galaxy ✍️ 🧐
- Introduction to Galaxy Analyses / A Short Introduction to Galaxy ✍️ 🧐
- Introduction to Galaxy Analyses / Options for using Galaxy 🧐
- Microbiome / Introduction to Microbiome Analysis ✍️ 🧐
- Microbiome / Introduction to metatranscriptomics ✍️ 🧐
- Teaching and Hosting Galaxy training / Overview of the Galaxy Training Material for Instructors 🧐
- Teaching and Hosting Galaxy training / Workshop Kickoff ✍️ 🧐
- Metabolomics / Introduction to Metabolomics 🧐
- Metabolomics / Predicting EI+ mass spectra with QCxMS 🧐
- Metabolomics / Mass spectrometry: LC-MS preprocessing - advanced 🧐
- Single Cell / Single-cell Formats and Resources 🧐
- Single Cell / Automated Cell Annotation 🧐
- Single Cell / An introduction to scRNA-seq data analysis 🧐
- Single Cell / GO Enrichment Analysis on Single-Cell RNA-Seq Data 🧐
- Single Cell / Trajectory analysis 🧐
- Single Cell / Plates, Batches, and Barcodes 🧐
- Climate / Pangeo ecosystem 101 for everyone 🧐
- Climate / Introduction to climate data 🧐
- Climate / Functionally Assembled Terrestrial Ecosystem Simulator (FATES) 🧐
- Climate / The Pangeo ecosystem 🧐
- Variant Analysis / Introduction to Variant analysis 🧐
- Sequence analysis / Quality Control 🧐
- Sequence analysis / Mapping ✍️ 🧐
- Visualisation / Visualisations in Galaxy ✍️ 🧐
- Visualisation / JBrowse ✍️ 🧐
- Visualisation / Friends Don't Let Friends Make Bad Graphs 🧐
- Visualisation / Circos ✍️ 🧐
- Galaxy Server administration / Ansible 🧐
- Galaxy Server administration / Galactic Database 🧐
- Galaxy Server administration / Galaxy Interactive Tools 🧐
- Galaxy Server administration / Galaxy from an administrator's point of view 🧐
- Galaxy Server administration / Reference Data with CVMFS 🧐
- Galaxy Server administration / Controlling Galaxy with systemd or Supervisor 🧐
- Galaxy Server administration / Advanced customisation of a Galaxy instance 🧐
- Galaxy Server administration / External Authentication 🧐
- Galaxy Server administration / Terraform 🧐
- Galaxy Server administration / uWSGI 🧐
- Galaxy Server administration / Gearing towards production 🧐
- Galaxy Server administration / Galaxy Tool Management with Ephemeris 🧐
- Galaxy Server administration / Server Maintenance: Cleanup, Backup, and Restoration 🧐
- Galaxy Server administration / Galaxy Installation with Ansible 🧐
- Galaxy Server administration / Reference Genomes in Galaxy 🧐
- Galaxy Server administration / User, Role, Group, Quota, and Authentication managment 🧐
- Galaxy Server administration / Galaxy on the Cloud 🧐
- Galaxy Server administration / Empathy ✍️ 🧐
- Galaxy Server administration / Galaxy Monitoring with Telegraf and Grafana 🧐
- Galaxy Server administration / Storage Management ✍️ 🧐
- Galaxy Server administration / Running Jobs on Remote Resources with Pulsar 🧐
- Galaxy Server administration / Connecting Galaxy to a compute cluster 🧐
- Galaxy Server administration / Galaxy Monitoring with gxadmin 🧐
- Galaxy Server administration / Storage Management 🧐
- Galaxy Server administration / Docker and Galaxy 🧐
- Galaxy Server administration / Galaxy Troubleshooting 🧐
- Galaxy Server administration / Server: Other 🧐
- Galaxy Server administration / Galaxy Monitoring 🧐
- Proteomics / Introduction to proteomics, protein identification, quantification and statistical modelling 🧐
- Using Galaxy and Managing your Data / Getting data into Galaxy ✍️ 🧐
- Using Galaxy and Managing your Data / Introduction to SRA Aligned Read Format and Cloud Metadata for SARS-CoV-2 🧐
- Using Galaxy and Managing your Data / Galaxy workflows in Dockstore 🧐
- Foundations of Data Science / Bioinformatics Data Types and Databases 🧐
- Foundations of Data Science / A brief history of modern biology 🧐
-
Imaging
/
Nucleoli Segmentation
&
Feature Extraction
using CellProfiler 🧐 - Transcriptomics / Introduction to Transcriptomics 🧐
- Transcriptomics / Integrate and query local datasets and distant RDF data with AskOmics using Semantic Web technologies 🧐
- Transcriptomics / Network Analysis with Heinz 🧐
- Transcriptomics / Visualization of RNA-Seq results with CummeRbund 🧐
- Epigenetics / EWAS Epigenome-Wide Association Studies Introduction 🧐
- Epigenetics / Introduction to DNA Methylation data analysis 🧐
- Epigenetics / Introduction to ChIP-Seq data analysis 🧐
- Epigenetics / ChIP-seq data analysis 🧐
- Epigenetics / Introduction to ATAC-Seq data analysis 🧐
- Contributing to the Galaxy Training Material / Overview of the Galaxy Training Material 🧐
- Contributing to the Galaxy Training Material / Contributing with GitHub via command-line 🧐
- Contributing to the Galaxy Training Material / Creating Slides ✍️ 🧐
- FAIR Data, Workflows, and Research / Intro to DataPLANT ARCs ✍️
- Statistics and machine learning / Gleam Image Learner - Validating Skin Lesion Classification on HAM10000 🧐
- Statistics and machine learning / Convolutional neural networks (CNN) Deep Learning - Part 3 🧐
- Statistics and machine learning / Gleam Multimodal Learner - HNSCC Recurrence Prediction with HANCOCK 🧐
- Statistics and machine learning / Recurrent neural networks (RNN) Deep Learning - Part 2 🧐
- Statistics and machine learning / Neural networks using Python 🧐
- Statistics and machine learning / Feedforward neural networks (FNN) Deep Learning - Part 1 🧐
- Statistics and machine learning / Deep Learning (without Generative Artificial Intelligence) using Python 🧐
- Statistics and machine learning / Regulations/standards for AI using DOME 🧐
- Assembly / Unicycler assembly of SARS-CoV-2 genome with preprocessing to remove human genome reads 🧐
- Assembly / An Introduction to Genome Assembly 🧐
- Assembly / Genome assembly quality control. 🧐
- Assembly / Deeper look into Genome Assembly algorithms 🧐
- Assembly / Genome assembly and assembly QC - Introduction short version 🧐
- Assembly / Unicycler Assembly 🧐
- Assembly / De Bruijn Graph Assembly 🧐
- Assembly / An introduction to get started in genome assembly and annotation 🧐
- Development in Galaxy / Galaxy Interactive Tours 🧐
- Development in Galaxy / Prerequisites for building software/conda packages 🧐
- Development in Galaxy / Introduction to the ToolFactory tutorial. 🧐
- Development in Galaxy / Tool Dependencies and Containers 🧐
- Development in Galaxy / Galaxy Webhooks 🧐
- Development in Galaxy / Tool development and integration into Galaxy ✍️ 🧐
- Development in Galaxy / Scripting Galaxy using the API and BioBlend 🧐
- Development in Galaxy / Tool Shed: sharing Galaxy tools 🧐
- Development in Galaxy / Generic plugins ✍️ 🧐
- Development in Galaxy / Visualizations: JavaScript Plugins ✍️ 🧐
- Development in Galaxy / Galaxy Interactive Environments ✍️ 🧐
- Development in Galaxy / Tool Dependencies and Conda 🧐
- Introduction to Galaxy Analyses / Una breve introducción a Galaxy ✍️ 🧐
- Single Cell / Introducción al análisis de datos de scRNA-seq 🧐
- Introduction to Galaxy Analyses / Una Breve Introducción a Galaxy ✍️ 🧐
- Single Cell / Una introducción al análisis de datos scRNA-seq 🧐
FAQs
- Can I create multiple Galaxy accounts?
- Log in to Galaxy using Single Sign-on
- How can I adapt this tutorial to my own data?
- How can I adapt this tutorial to my own data?
- Adding a custom database/build (dbkey)
- How can I do analysis X? - Getting help
- My jobs aren't running!
- Regular Expressions 101
- Ergebnisse können variieren ✍️
- I risultati possono variare ✍️
- Results may vary
- Los resultados pueden variar ✍️
- Troubleshooting errors
- Define once, reference many times
- Adding a tag to a collection
- Erstellen einer Datensatzsammlung ✍️
- Creare una raccolta di set di dati ✍️
- Creating a dataset collection
- Crear una colección de conjuntos de datos ✍️
- Erstellen einer gepaarten Sammlung ✍️
- Creazione di una raccolta accoppiata ✍️
- Creating a paired collection
- Creación de una colección emparejada ✍️
- Changing the datatype of a collection
- Converting the datatype of a collection
- Renaming a collection
- How do I find the Community Home pages?
- Thanks!
- How can I get started with contributing?
- How can I contribute in "advanced" mode?
- How can I give feedback?
- What can I do to help the project?
- How can I report mistakes or errors?
- How can I test an Interactive Tour?
- How can I create new content without dealing with git?
- Hinzufügen eines Tags ✍️
- Aggiunta di un tag ✍️
- Adding a tag
- Añadir una etiqueta ✍️
- Ändern des Datentyps ✍️
- Modifica del tipo di dato ✍️
- Changing the datatype
- Changing database/build (dbkey)
- Converting the file format
- Erstellen einer neuen Datei ✍️
- Creare un nuovo file ✍️
- Creating a new file
- Creación de un nuevo fichero ✍️
- Detecting the datatype (file format)
- Importieren von Daten aus einer Datenbibliothek ✍️
- Importare i dati da una libreria di dati ✍️
- Importing data from a data library
- Importar datos de una biblioteca de datos ✍️
- Importing data from repositories
- Importieren über Links ✍️
- Importazione tramite link ✍️
- Importing via links
- Umbenennen eines Datensatzes ✍️
- Rinominare un set di dati ✍️
- Renaming a dataset
- Cambiar el nombre de un conjunto de datos ✍️
- Upload many files (>10) via FTP
- Compressed FASTQ files, (`*.gz`)
- FASTQ files: `fastq` vs `fastqsanger` vs ..
- Verwendung des Fenstermanagers zur Anzeige mehrerer Datensätze ✍️
- Usare la Gestione finestre per visualizzare più insiemi di dati ✍️
- Using the Window Manager to view multiple datasets
- Where can I read more about Quality Control of data?
- How many mules?
- Are there any upcoming events focused on Galaxy Training?
- Switching Galaxy Labs
- What is Galaxy?
- Forking the GTN repository
- Updating the default branch from master to main
- Syncing your Fork of the GTN
- GTN ADR: Image Storage
- GTN ADR: Why Jekyll and not another Static Site Generator (SSG)
- GTN Architectural Decision Record Template
- Using tutorial mode and the Case Study suite
- What is this website?
- How can I advertise the training materials on my posters?
- What audiences are the tutorials for?
- How can I cite the GTN?
- Creating a GTN Event
- How is the content licensed?
- Creating a GTN News post
- Sustainability of the training-material and metadata
- Supporting Tutorial Mode (GTN-in-Galaxy) in a tutorial
- What are the tutorials for?
- Kopieren eines Datensatzes zwischen Historien ✍️
- Copiare un set di dati tra le cronologie ✍️
- Copy a dataset between histories
- Copiar un conjunto de datos entre historiales ✍️
- Erstellen eines neuen Verlaufs ✍️
- Creare una nuova cronologia ✍️
- Creating a new history
- Para la creación de un historial nuevo ✍️
- Créer un nouvel history
- Exporting a history
- Export tool citations from a history
- Finding Histories
- Umbenennen eines Verlaufs ✍️
- Rinominare una cronologia ✍️
- Renaming a history
- Searching your history
- Sharing your History
- Which icons are available to use in my tutorial?
- The input for a tool is not listed in the dropdown
- The input for a tool is not listed in the dropdown
- Install tools via the Admin UI
- What are the best practices for teaching with Galaxy?
- What Galaxy instance should I use for my training?
- How do I get help?
- Where do I start?
- Launch JupyterLab
- Open interactive tool
- Launch RStudio
- Stop RStudio
- Stop an Interactive Tool
- JBrowse is taking a long time to complete?
- How can I get help?
- Where do I start?
- Where can I run the hands-on tutorials?
- How do I use this material?
- Defining a Learning Pathway
- How do I find the Maintainer Home pages?
- Where can I read more about this analysis?
- Where can I read more about this analysis?
- MultiQC error for your FastQC reports?
- I cannot run client tests because yarn is not installed.
- Getting your API key
- Adding your recording to a tutorial or slide deck
- Recording a video tutorial
- Illumina MiSeq sequencing
- Nanopore sequencing
- What is a SNP?
- Where do I get more support?
- Contacting Galaxy Administrators
- Ändern der Werkzeugversion ✍️
- Cambiare la versione dello strumento ✍️
- Changing the tool version
- Cambiar la versión de la herramienta ✍️
- Organizing the tool panel
- Viewing tool logs (`stdout` and `stderr`)
- If a Tool is Missing
- Re-running a tool
- Auswählen einer Datensatzsammlung als Eingabe ✍️
- Selezione di una raccolta di dati come input ✍️
- Selecting a dataset collection as input
- Selección de una colección de conjuntos de datos como entrada ✍️
- Mehrere Datensätze auswählen ✍️
- Selezionare più insiemi di dati ✍️
- Select multiple datasets
- Seleccionar varios conjuntos de datos ✍️
- Multipile ähnliche Werkzeuge verfügbar ✍️
- Strumenti simili alla compilazione multipla disponibili ✍️
- Multipile similar tools available
- Múltiples herramientas similares disponibles ✍️
- How can I create a tutorial skeleton from a Galaxy workflow?
- Tutorial-Modus verwenden ✍️
- Utilizzo della modalità tutorial ✍️
- Using tutorial mode
- Utilizar modo tutorial ✍️
- Using IGV with Galaxy
- Add genome and annotations to IGV from Galaxy
- Add Mapped reads track to IGV from Galaxy
- What is a Learning Pathway?
- Creating a new workflow
- Downloading workflows
- Opening the workflow editor
- Extracting a workflow from your history
- Hiding intermediate steps
- Importing a workflow
- Setting parameters at run-time
- Renaming workflow outputs
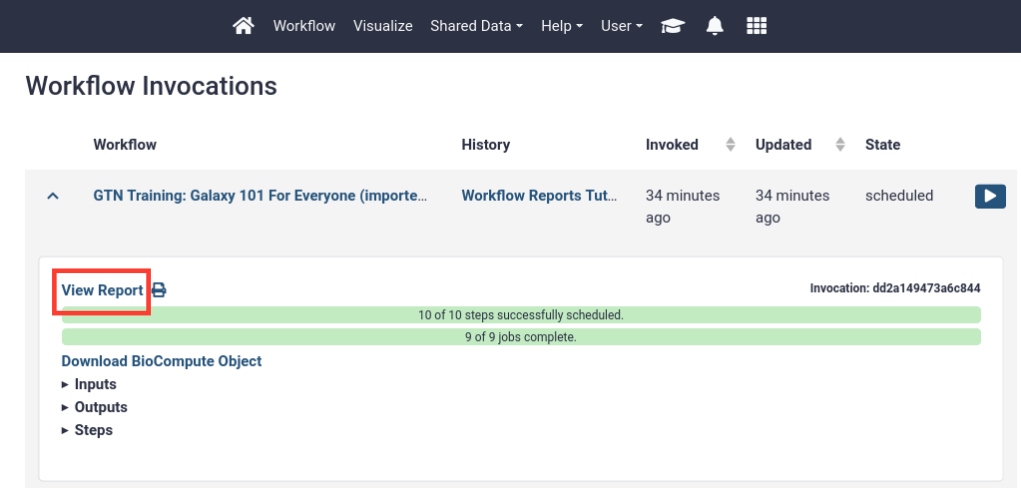
- Viewing a workflow report
- Running a workflow
- Share
Video Recordings
- Microbiome / Introduction to Microbiome Analysis 💬 🗣
- Visualisation / Circos 💬
- Galaxy Server administration / Running Jobs on Remote Resources with Pulsar 💬
- Transcriptomics / Introduction to Transcriptomics 💬
- Contributing to the Galaxy Training Material / Creating Slides 💬
- Assembly / An Introduction to Genome Assembly 💬
- Introduction to Galaxy Analyses / Galaxy Basics for genomics 💬
- Microbiome / 16S Microbial Analysis with mothur (short) 💬 🗣
- Microbiome / Metatranscriptomics analysis using microbiome RNA-seq data 💬 🗣
- Microbiome / Metatranscriptomics analysis using microbiome RNA-seq data (short) 💬 🗣
- Sequence analysis / Quality Control 💬
- Sequence analysis / Mapping 💬
- Visualisation / Visualisation with Circos 💬
- Galaxy Server administration / Mapping Jobs to Destinations using TPV 💬
- Galaxy Server administration / Galaxy Installation with Ansible 💬
- Galaxy Server administration / Galaxy Monitoring with Telegraf and Grafana 💬
- Galaxy Server administration / Data Libraries 💬 🗣
- Galaxy Server administration / Connecting Galaxy to a compute cluster 💬
- Galaxy Server administration / Training Infrastructure as a Service (TIaaS) 💬
- Using Galaxy and Managing your Data / Workflow Reports 💬 🗣
- Using Galaxy and Managing your Data / Understanding Galaxy history system 💬 🗣
- Transcriptomics / Reference-based RNA-Seq data analysis 💬
- Transcriptomics / RNA Seq Counts to Viz in R 💬
- Contributing to the Galaxy Training Material / Preview the GTN website as you edit your training material 💬 🗣
- Sequence analysis / Qualitätskontrolle 💬
- Sequence analysis / Mapping 💬
- Transcriptomics / Referenzbasierte RNA-Seq-Datenanalyse 💬
- Sequence analysis / Control de calidad 💬
- Sequence analysis / Mapeo 💬
- Transcriptomics / Análisis de datos RNA-Seq basados en referencias 💬
- Sequence analysis / Controllo qualità 💬
- Sequence analysis / Mappatura 💬
- Transcriptomics / Analisi dei dati RNA-Seq basata su riferimenti 💬
Events
- Admin Training @ GCC 2022: Online Training Day 🎪
- My External Training Event Title 🎪
- My Training Event Title 🎪
- Galaxy Training Academy 2025 🎪
- LOVE DATA week with intro to Galaxy & GTN 🎪
- Galaxy Training Academy 2026 🎪
- Galaxy Training Academy 2024 🎪
GitHub Activity
github Issues Reported
412 Merged Pull Requests
See all of the github Pull Requests and github Commits by Saskia Hiltemann.
-
 [Infra] Add link to youtube cover images to video processing checklist
template-and-tools
[Infra] Add link to youtube cover images to video processing checklist
template-and-tools -
 Add available translations to the tutorial overview box
template-and-tools
Add available translations to the tutorial overview box
template-and-tools -
 Fix deploy error
template-and-tools
Fix deploy error
template-and-tools -
 [Intro Galaxy RDM] Tweak tutorial
introductiontemplate-and-toolsfaqs
[Intro Galaxy RDM] Tweak tutorial
introductiontemplate-and-toolsfaqs -
 [New Tutorial] Intro to Galaxy as an RDM Platform
introductiontemplate-and-toolsfaqs
[New Tutorial] Intro to Galaxy as an RDM Platform
introductiontemplate-and-toolsfaqs
Reviewed 1481 PRs
We love our community reviewing each other's work!
-
 Update recentrifuge input in bacterial isolate contamination tutorial
ecology
Update recentrifuge input in bacterial isolate contamination tutorial
ecology -
 Bump addressable from 2.8.5 to 2.9.0
template-and-toolsdependenciesruby
Bump addressable from 2.8.5 to 2.9.0
template-and-toolsdependenciesruby -
 Update 2026-10-12-Advanced-Galaxy-Training : add requirements, keep only FR server
template-and-tools
Update 2026-10-12-Advanced-Galaxy-Training : add requirements, keep only FR server
template-and-tools -
 correcting recent news item - Fix links and formatting in biont.md
news
correcting recent news item - Fix links and formatting in biont.md
news -
 [News Item] Add BioNT project announcement for translated tutorials
news
[News Item] Add BioNT project announcement for translated tutorials
news
News

Next GTN CoFest May 20, 2021
New Feature: FAQs

New Tutorials: Genome assembly of a MRSA genome

¿Hablas español?: The first curated tutorial in Spanish!

Oh no, it changed! Quick, to the archive menu.
New Tutorial: GitPod for contributing to the GTN

New Tutorial: Data Manipulation
GTN Video Library 2.0: 107 hours of learning across 154 videos

GTN Video Library 2.0: 107 hours of learning across 154 videos

4th Mycobacterium tuberculosis complex NGS made easy

An Ode to Helena - from the 🥓Bacon Brigade

GTN is now integrated with WorkflowHub
External Links

Favourite Topics
Favourite Formats